Automated Graphene Clustering
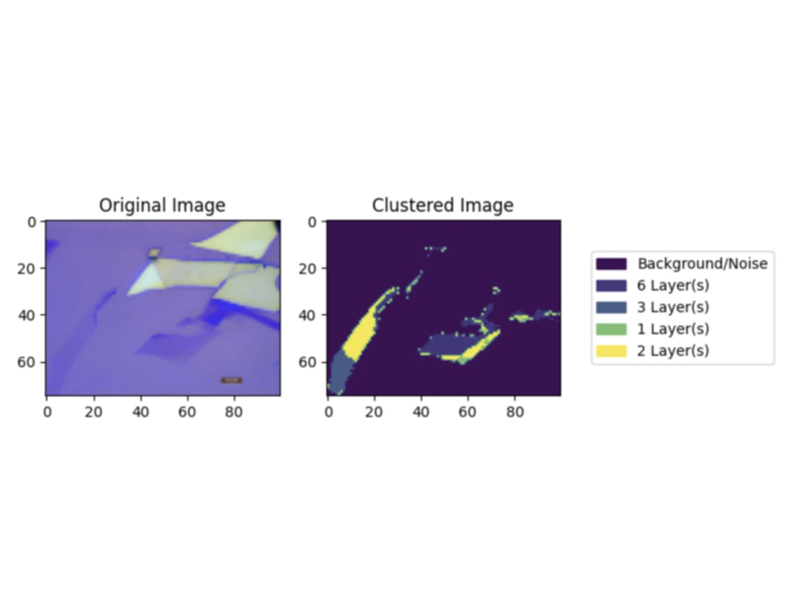
I designed a pipeline that took an image of a Graphene sample on an \({\rm Si}/{\rm SiO}_2\) substrate and returned a color-coded image that identifies the layers of the sample on the substrate. The pipeline took an image and first passed it through a binary convolutional neural network (CNN) to identify if a sample was present on the image in the first place, to reduce processing overhead. Then, after pre-processing the images using techniques from similar papers, like noise cleaning, edge detection Canny methods, and Gaussian blurring, we pass the images into multiple unsupervised clustering techniques. We utilize DBSCAN and GMM to cluster the image, identifying the layers without a ground truth, and then returned the clustered image with a key. We then built a simple user interface on top of this pipeline in order to be able to use the program.
Below is the poster presentation given on this project:
Behind the Model
We created a convolutional neural network (CNN) to identify if a sample is present on a given image.
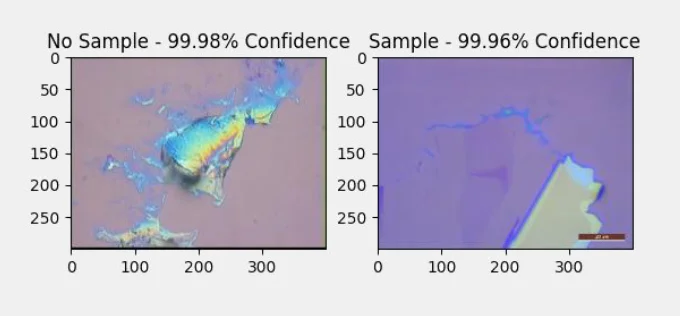
Sample identification with confidence of an image with and without a sample.
We them pre-process the image. We downsize the image to reduce features and save time iterating through every pixel of an image. We then covert from RGB colorspace to LAB color space, which we use to extract the background image. We then use a median blur method to smooth out noise in the image. We then isolate the L values to use for Canny edge detection, locating the contours and isolating the sample from the substrate.
We then put the isolated image through DBSCAN, which finds core samples in high density regions and expands clusters from them. However, clusters classified in DBSCAN are more than are actually present due to samples of the same class being separated in distance. To solve this, we employed Gaussian Mixture Modeling (GMM), which helps find similarities between clusters. So, from a sample set of size \(N\), GMM predicts \(K\) clusters, which the user defines.
After the image is processed by the GMM model, we have to clean the image by reclassifying stray pixels (classified pixels that weren’t identified as being within the sample edges) and far pixels (normalized value in RGB colorspace deviates greater than 5% compared to all other pixels of its class).
For more information on the model, the mathematics, or the results, see the poster above.
Acknowledgements
This project acknowledges the following:
- Jacob Balek: For work on a similar project throughout the summer with a differing methodology.
- Jiayin Wang and Dr. ChunNing Lau: For support, guidance, and mentorship in the laboratory setting.
- Max Sullivan: For code testing and data collection.
- Xuanzhi Zhang: For insight into a similar project completed for an undergraduate thesis.
- Greyson Voigt: For access to Graphene data files.
- Lindsey Thaler/The Ohio State University Department of Physics/Bunny C. Clark Student Support Fund: For funds, support, and setting up of this project through the 2023 OSU Physics Summer Research Program.
References
Masubuchi, S., Morimoto, M., Morikawa, S., Onodera, M., Asakawa, Y., Watanabe, K., Taniguchi, T., and Machida, T. (2018). Autonomous robotic searching and assembly of two-dimensional crystals to build van der waals superlattices. Nature Communications, 9(1413).
Sterbentz, R. M., Haley, K. L., and Island, J. O. (2021). Universal image segmentation for optical identification of 2d materials. Scientific Reports, 11(5808).
Wang, Y. Y., Gao, R. X., Ni, Z. H., He, H., Guo, S. P., Yang, H. P., Cong, C. X., and Yu, T. (2012). Thickness identification of two-dimensional materials by optical imaging. Nanotechnology, 23(49):495713.
Zhang, X. (2023). Similar color segmentation using number of cluster algorithm.
